Welcome
Welcome to the Molecular Signatures with Biclustering (MoSBi) webtool!
We apply biclustering algorithms on omics data in combination with a novel ensemble approach and find to find molecular signatures in the data. This can yield to new insights of the data, which go further than traditional clustering, where only one dimension is considered.
MoSBi is not only available as a public webservice, but can also be used as an R package, or the webserver can be deployed on your own computer using Docker. To download and run MoSBi locally, look at the
Aboutpage.
Before using MoSBi, we recommend reading the publication:
Rose et al. "MoSBi: Automated signature mining for molecular stratification and subtyping"
Proceedings of the National Academy of Sciences
, 2022; 119 (16): e2118210119;
doi:
https://doi.org/10.1073/pnas.2118210119
Please note that your uploaded data will be processed on our servers and automatically be deleted once your session is closed. Note we have only limited capacities on our server. If page freezes occur or your data exceeds the limit, consider running MoSBi locally. Further information are available on the
Aboutpage.
1. Upload Data
Upload your data in the format as explained on the 'Tutorial' page. You can additionally upload labels and colors for your data. Once data is uploaded, you can proceed to the computation tab.
The data size is limited to 1e+07 rows and 1e+07 columns. Download and run MoSBi on your machine, if your data exceeds this limit. For more information visit the 'About' page.
Example data from:
Ku et al., Molecular Oncology (2020)Data summary
UMAP visualization
2. Compute Biclusters
2.1. Algorithms for bicluster computation
Please select one of multiple biclustering algorithms that will be run with your uploaded data. The results of all algorithms will be combined for the ensemble analysis.
The preselected algorithms were selected due to runtime and performance. Other algorithms can result in extensive runtimes. Inexperienced users should not change the selection.
2.2. Optional Parameters for the biclustering algorithms
Spectral
Biclustpy
Bi-Force
akmbiclust
2.3. Run Algorithms
Please press 'Compute Biclusters' before proceeding to the next page.
3. Biclustering Results
3.1. Distribution of bicluster sizes
Gives an overview of the sizes of biclusters.
3.2. Biclusters
Heatmaps of all computed biclusters, sorted by algorithm.
3.3. Download biclusters
Biclusters can be downloaded here, or later on page 4.
Download Biclusters (as xlsx table)4. Molecular Signatures
4.1. Ensemble parameters (Optional)
4.2. Calculate Networks
After the bicluster calculations, bicluster networks and communities can be computed here:
4.3. Error model statistics
Visualization of the cut-off estimation with our error model.
4.4. Bicluster networks
Colored by conditions
Colored by algorithm
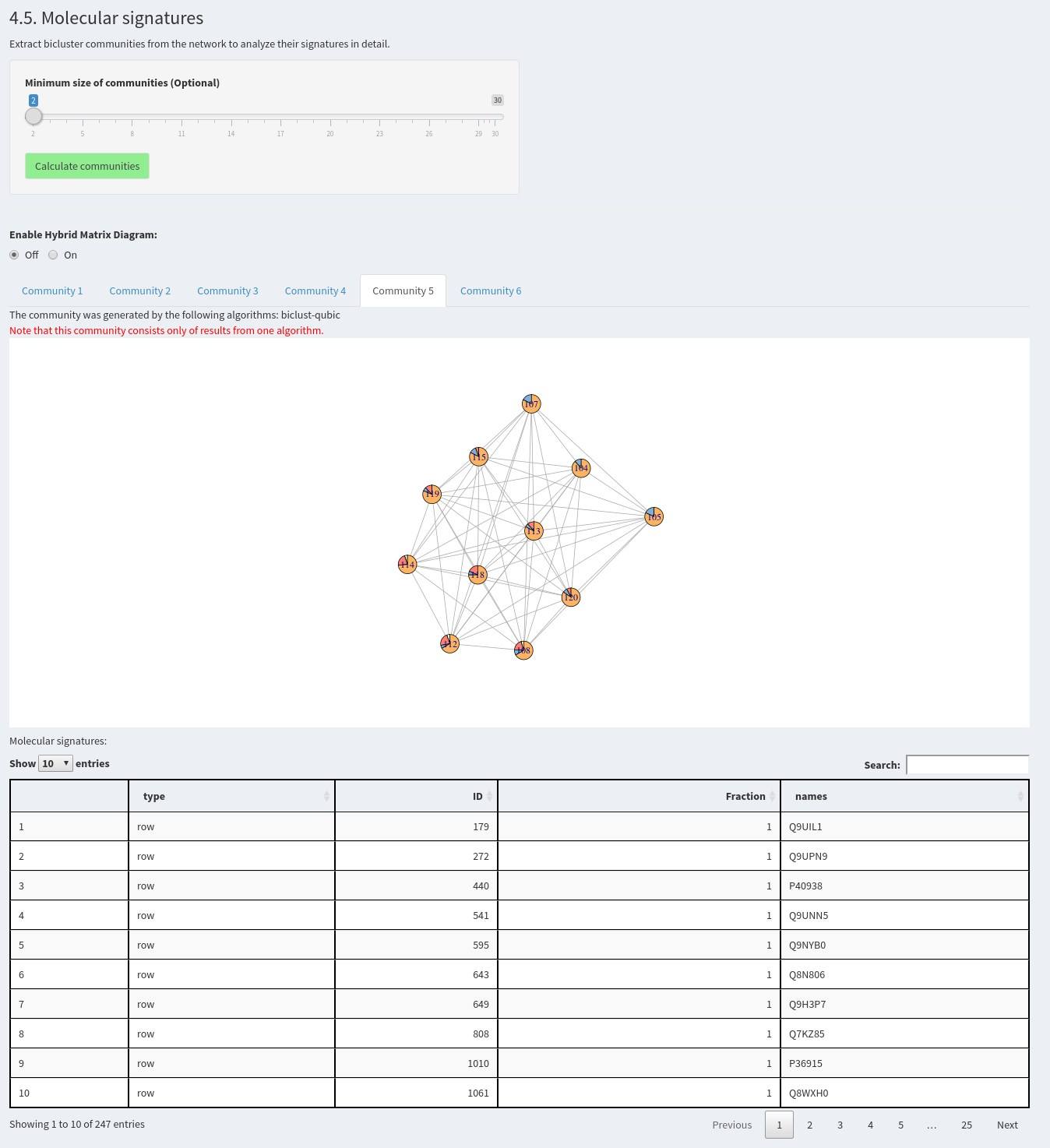
4.5. Molecular signatures
Extract bicluster communities from the network to analyze their signatures in detail.
4.6. Download Results
All results can be downloaded here as a zip file.
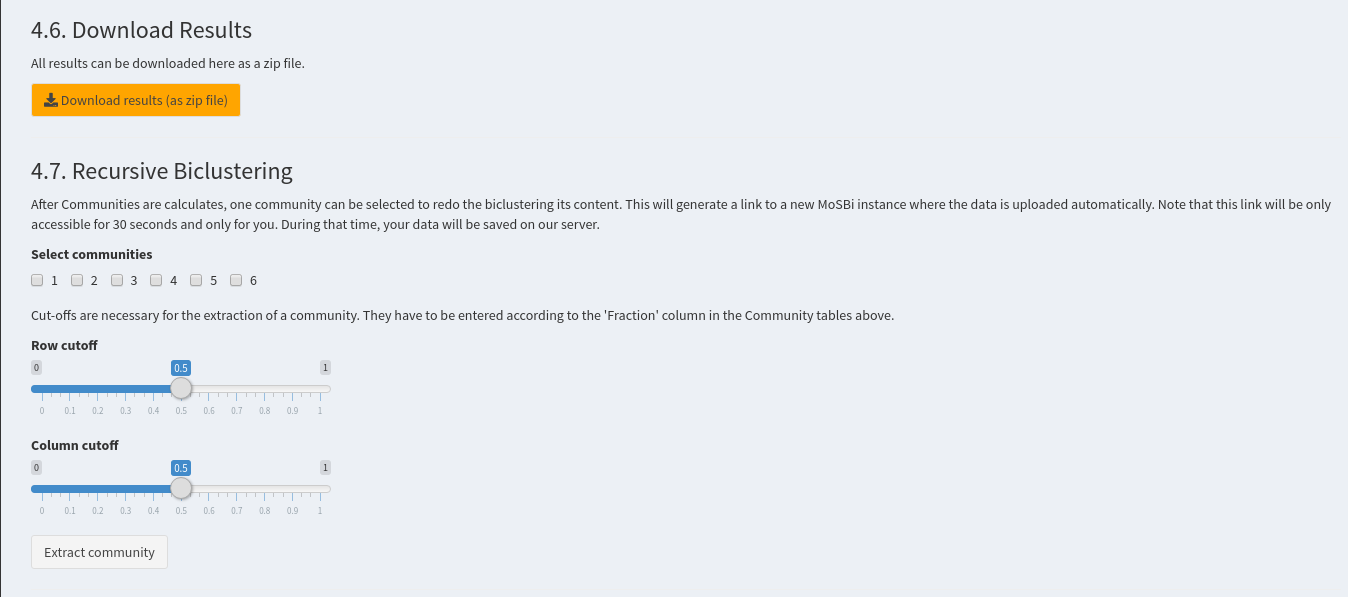
Download results (as zip file)(4.7. Recursive Biclustering)
After Communities are calculates, one community can be selected to redo the biclustering its content. This will generate a link to a new MoSBi instance where the data is uploaded automatically. Note that this link will be only accessible for 30 seconds and only for you. During that time, your data will be saved on our server.
Cut-offs are necessary for the extraction of a community. They have to be entered according to the 'Fraction' column in the Community tables above.
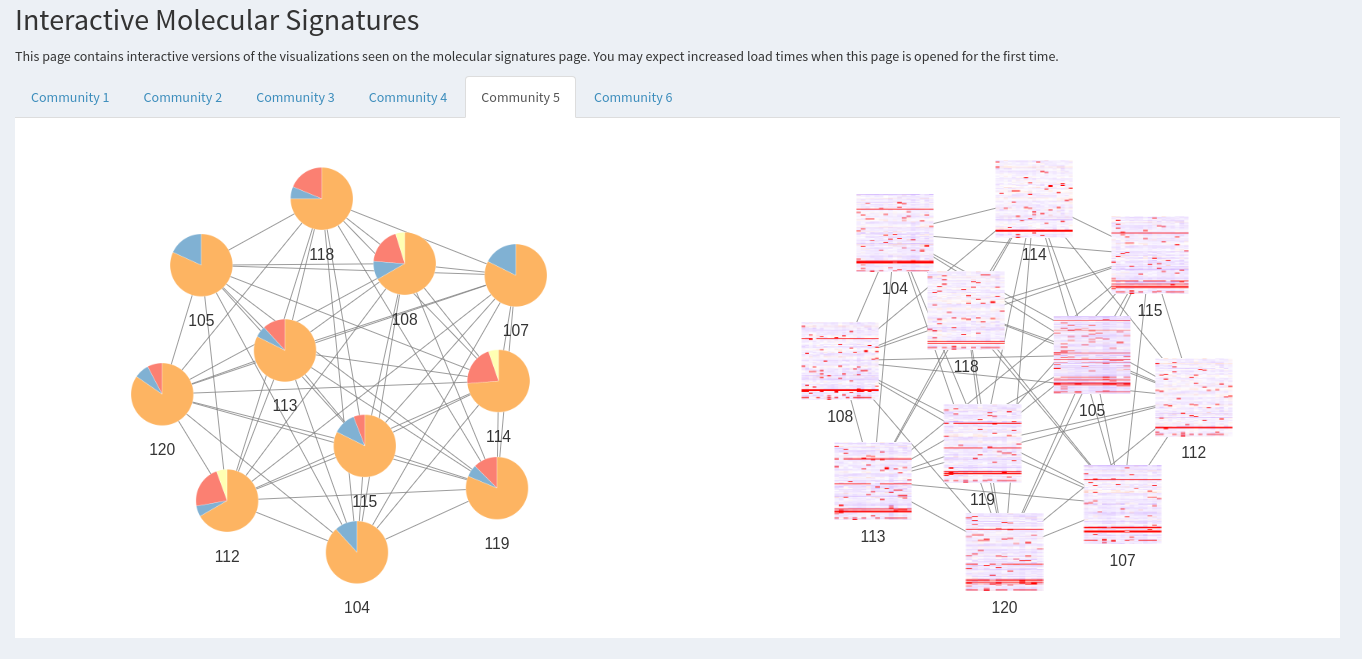
Interactive Molecular Signatures
This page contains interactive versions of the visualizations seen on the molecular signatures page. You may expect increased load times when this page is opened for the first time.
Tutorial
This tutorial will walk you through the main functionalities of the webtool. For a full analysis all steps 1.-4. need to be completed.
1. Data upload
To start the analysis, a dataset must be uploaded. An example dataset can be used by clicking on the button
Use example dataor user data can be uploaded. Additionally, labels for the samples in the dataset can be uploaded and custom colors which will be used in the generated plots. However, the latter two are optional and can be added at any time, even after the analysis is done and the plots will be updated automatically with the new colors or conditions.
The data must be uploaded in
.csvformat (Comma separated values). Here are examples how the upladed data should look like:
Dataset (required)
| Names | Sample1 | Sample2 | ... |
|---|---|---|---|
| Gene1 | 0.12 | 2.35 | ... |
| Gene2 | 5.12 | 0.98 | ... |
| ... | ... | ... | ... |
Labels for samples (Optional)
| id | condition |
|---|---|
| Sample1 | Disease1 |
| Sample2 | Disease2 |
| ... | ... |
Colors (Optional)
| condition | color |
|---|---|
| Disease1 | green |
| Disease2 | red |
| ... | ... |
Row and column names are required for the dataset. If no labels are uploaded, they can also be generated using k-means clustering. Other options include the possibility to normalize the data and replace missing values. Once the data is uploaded, the size of the dataset is shown with the number of missing values and a UMAP visualization is presented to give an overview of the data and labels.
The data size is limited on our servers. Therefore the maximum number of rows is limited to 1e+07 and the number of columns is limited to 1e+07. If your data exceeds that limit please run MoSBi locally. Information for that can be found on the About page.
The data is now ready to continue with the analysis and compute biclusters.
2. Compute biclusters
The first step of the analysis is the computation of biclusters, using multiple algorithms. Without prior experience or knowledge of biclustering, is recomended to use the prior selection of algorithms. Of you are only interested in the result of one algorithm, it is also sufficient to select only one algorithm.
The computation can now be started by clicking the
Compute biclustersbutton. This will execute all algorithms with default parameters. Parameters of all algorithms can be changed in the blue boxes for each parameter individually. A box in the lower right corner indicates the progress of the computation:
Once the computation has finished, an overview about the executed algorithms is shown, indicating if errors occured in some algorithms with a list of all error messages, which allows the you to adapt algorithm parameters and rerun the algorithms.
In the next step, the resulting biclusters can be investigated.
3. Biclustering Results
This page gives you an overview about the computed biclusters and the option to download them.
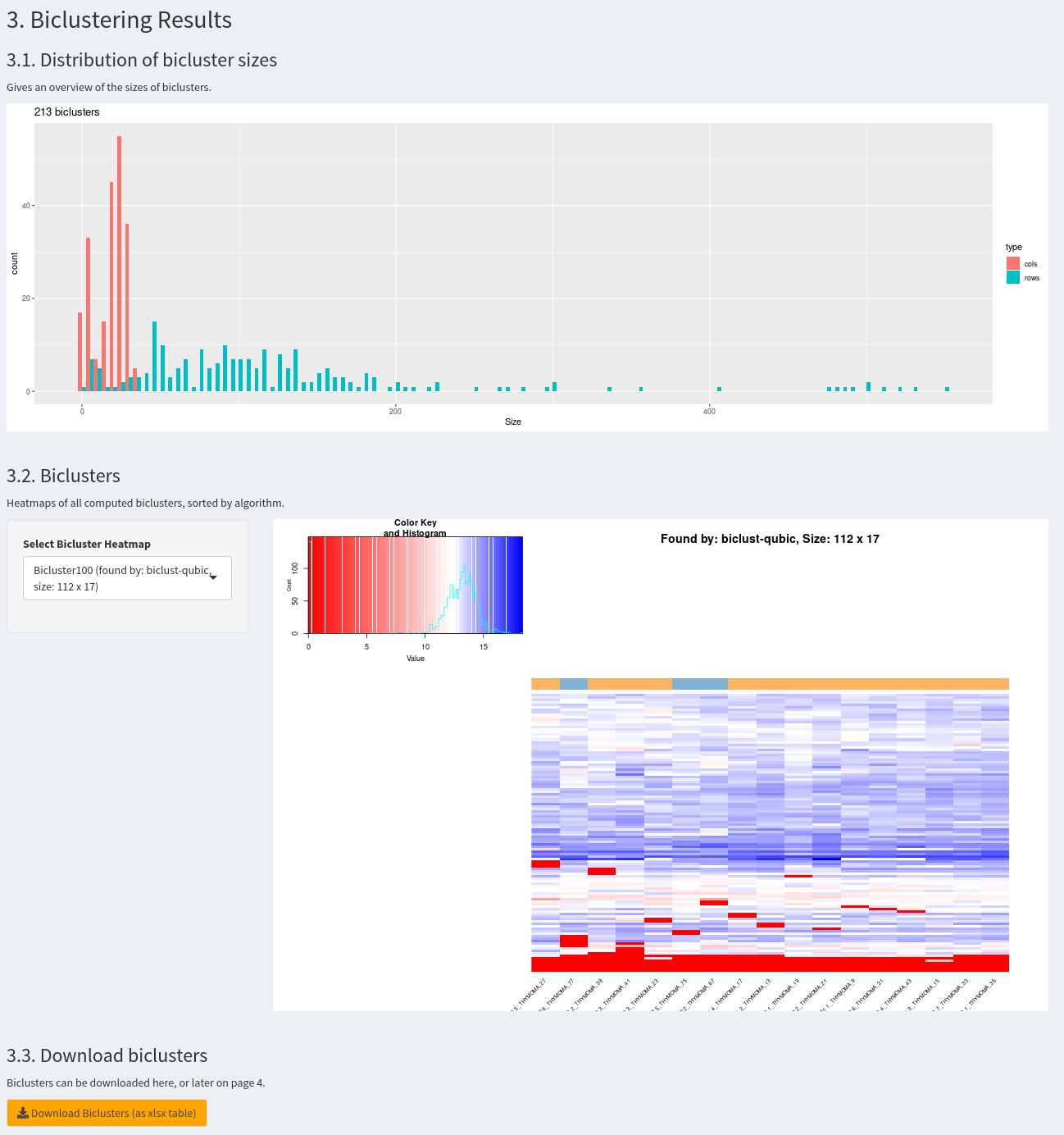
The histogram 3.1. plots the size distribution of all biclusters for rows and columns. Additionally, in 3.2. each bicluster can be selected individually and a heatmap with labels for samples can be inspected. Finally, All biclusters can be downloaded as an xlsx table. Here is an example how a resulting page can look like:
4. Molecular Signatures
The final step is the extraction of molecular signatures by creating bicluster networks and community detection.
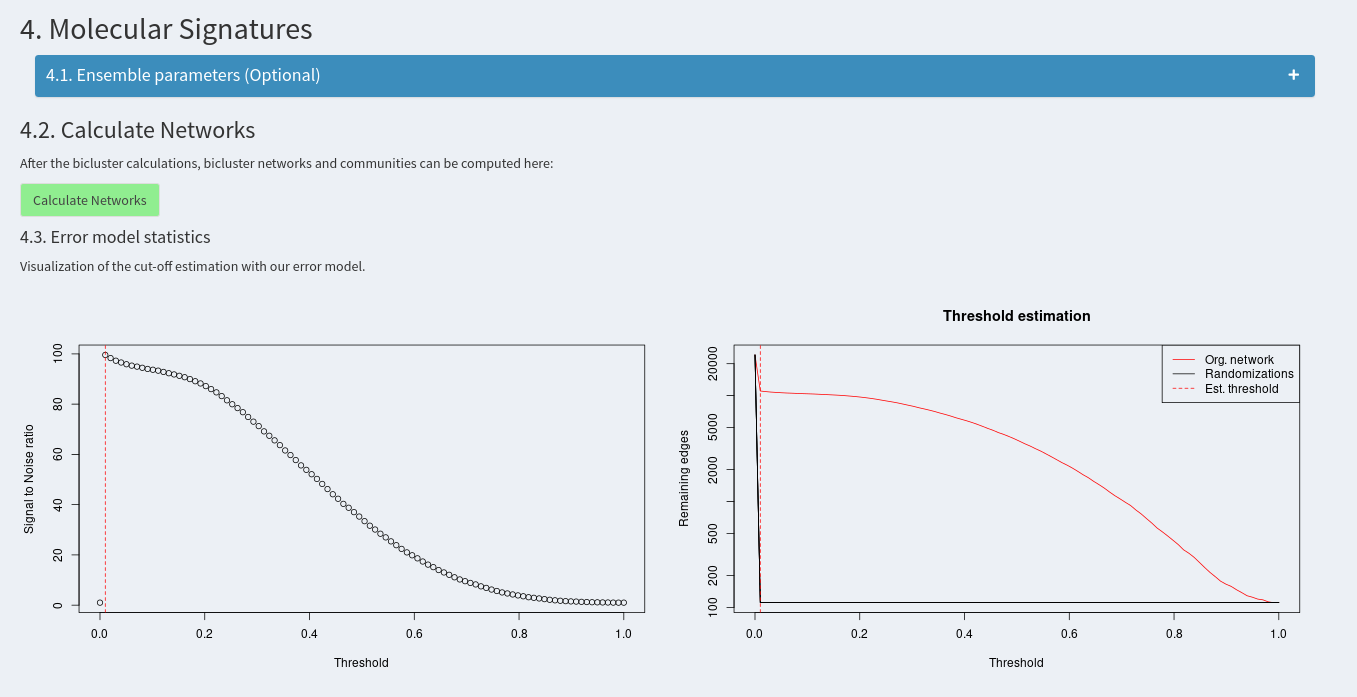
Additional parameters can be modified, however this is not recommended and should only be done after reading the publication. The networks can then be calculated using the accoding green button. The first two plots then show the estimation of the similarity threshold. For further information, please consult the publication.
The next plots shows the bicluster similarity networks colored by labels for the samples and by biclustering algorithm. Two connected biclusters show a higher than random similarity. On the left, the biclusters are colored by the sample labels that are included in the biclusters. This gives you an overview about the network and the similarities in the results of different algorithms.
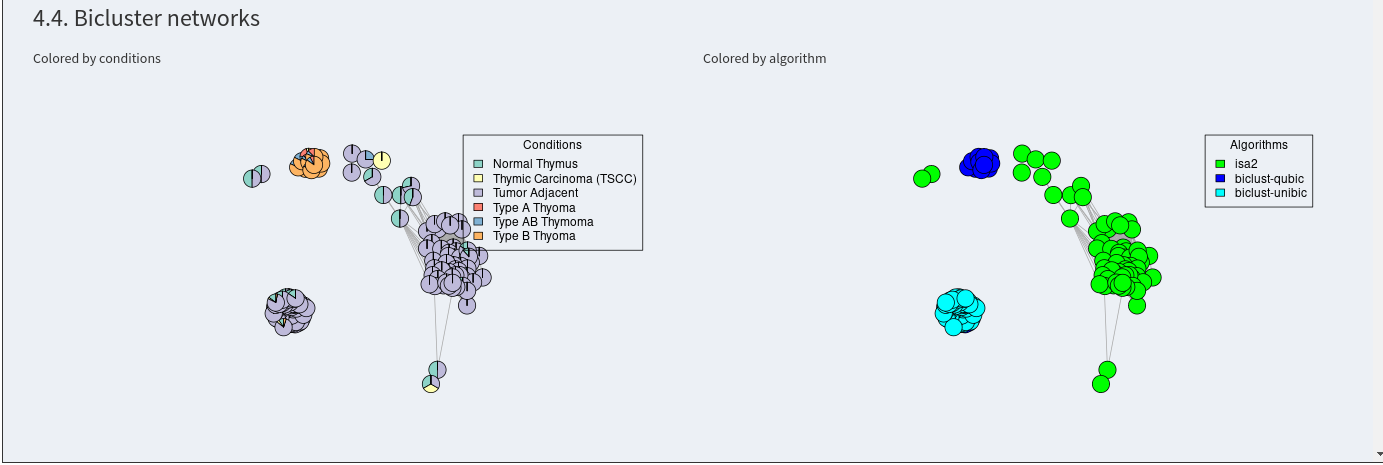
Now, communities can be calculated to extract the signatures. A minimum community size can be defined. Afterwards, all communities will be shown in tabs below as networks and tables, where the included samples and features can be investigated. The visualization of the network as a hybrid matrix diagram plots bicusters as heatmatps in the network. This helps to visually inspect the similarity of biclusters.
All results can be downloaded as a zip file afterwards. Additionally, it is possible to extract the content of a community and perform another round of biclustering on this subsetted data. This will temporarily save the date on our server and make it available through the public link that occurs after clicking on the according button. The link will be invalidated after 30 seconds. By clicking again a new link is generated.
The page
Interactive Molecular Signaturesprovides the same community network plots, but in an interactive form.
About
Thank you for using MosBi!
Citation
If you use MoSBi, please cite our publication:
Rose et al. "MoSBi: Automated signature mining for molecular stratification and subtyping"
Proceedings of the National Academy of Sciences
, 2022; 119 (16): e2118210119;
doi:
https://doi.org/10.1073/pnas.2118210119
Feedback
If you have questions or suggestions how to improve MoSBi, please contact us:
<tim.rose[a.t_)tum.de>
<josch.pauling[a.t_)tum.de>
Code
MoSBi is free software. If you want to run MoSBi locally or have a look at the code here:
Impressum
Dr. Josch K. Pauling
bidt Junior Research Group LipiTUM
Chair of Experimental Bioinformatics
TUM School of Life Sciences Weihenstephan
Maximus-von-Imhof-Forum 3
85354 Freising
Germany